Information
- Ph.D., Oregon Health Sciences University, 2000
One of the major unsolved mysteries in molecular biology is the protein folding problem,
how proteins adopt their three-dimensional, biologically active conformations. In
particular, we are interested in understanding the mechanisms responsible for the
formation of membrane spanning channels that transport proteins important for disease
pathogenesis. Our laboratory focuses on two biological systems– the anthrax protective
antigen from Bacillus anthracis, and CooC, a protein required for transport of CS1
pili from enterotoxigenic E. coli to the bacterial cell surface. We utilize a variety
of biochemical and biophysical techniques, including circular dichroism, fluorescence
and NMR to aid in our investigations.
The anthrax protective antigen is the B component of the anthrax toxin, an AB toxin
secreted by B. anthracis that includes the enzymatic A components edema factor and
lethal factor. Edema and lethal factor enter the cell cytosol through a pore formed
by the protective antigen. The exact mechanism of pore formation is not understood,
but is known to require the pH-dependent release of a receptor (anthrax toxin receptors
1 or 2) from within an endocytic vesicle. Understanding what initiates the release
of the receptor and subsequent membrane insertion is of particular interest to our
laboratory. Our laboratory has developed methods for biosynthetically incorporating
2- and 4-fluorohistidine into proteins. 2-fluorohistidine has a side-chain pKa of
1, and uniform labeling of the protective antigen with this amino acid has been used
to address the role of histidine protonation in governing the conformational change
from a prepore to a membrane spanning pore, as well as receptor release.
CS1 pili from enterotoxigenic E. coli are long hair-like fibers that extend from the
bacterial surface and are required for adherence to host cells. The CS1 pilus assembly
system utilizes only four proteins for proper assembly of a pilus – CooA, the major
pilus subunit that comprises the polymeric fiber; CooB, a chaperone required for folding
of CooA; CooC, an outer membrane usher protein required for assembly at the outer
membrane; and CooD, the tip of the pilus which contains a lectin domain for interacting
with host cell surfaces. How pili, including CS1, are assembled is not well understood.
Of central importance is CooC, the membrane spanning usher protein required for transport
of CS1 pili to the cell surface. Through our studies on the related Pap system from
pylenophritic E. coli, CooC initiates the transfer of a chaperone subunit complex
(CooB-CooA), to a subunit-subunit complex (CooA-CooA). Understanding how CooC facilitates
this transfer process is a major focus of our research. Understanding the mechanism
of CS1 pilus assembly and transport at the outer membrane may aid in the development
of therapeutics that block pilus assembly or transport.
The anthrax protective antigen (PA) is an 83 kDa protein secreted by Bacillus anthracis, the causative agent of anthrax disease. Upon infection by B anthracis, PA and two other proteins, edema factor (EF) and lethal factor (LF) are secreted from the bacterium into the host bloodstream. PA binds to host cellular receptors, and after proteolytic processing assembles into heptameric (Lacy et al., PNAS 2004) or octameric (Kintzer et al., JMB 2009) ring-shaped structures called pre-pores. These pre-pore structures allow EF and LF to bind, and the entire complex is endocytosed into the host cell. Eventually, the resulting endosome becomes acidified as it is trafficked along, and forms a membrane spanning pore. The structure of the pore has recently been visualized in stunning detail using cryo-EM (Jiang et al., Nature 2015). Our research is directed towards understanding the mechanistic details of how PA forms a membrane spanning pore at low pH. Formation of the pore is absolutely critical for the pathogenesis of the toxin, since the pore provides a conduit for entry of the edema or lethal factor into the cell cytosol.
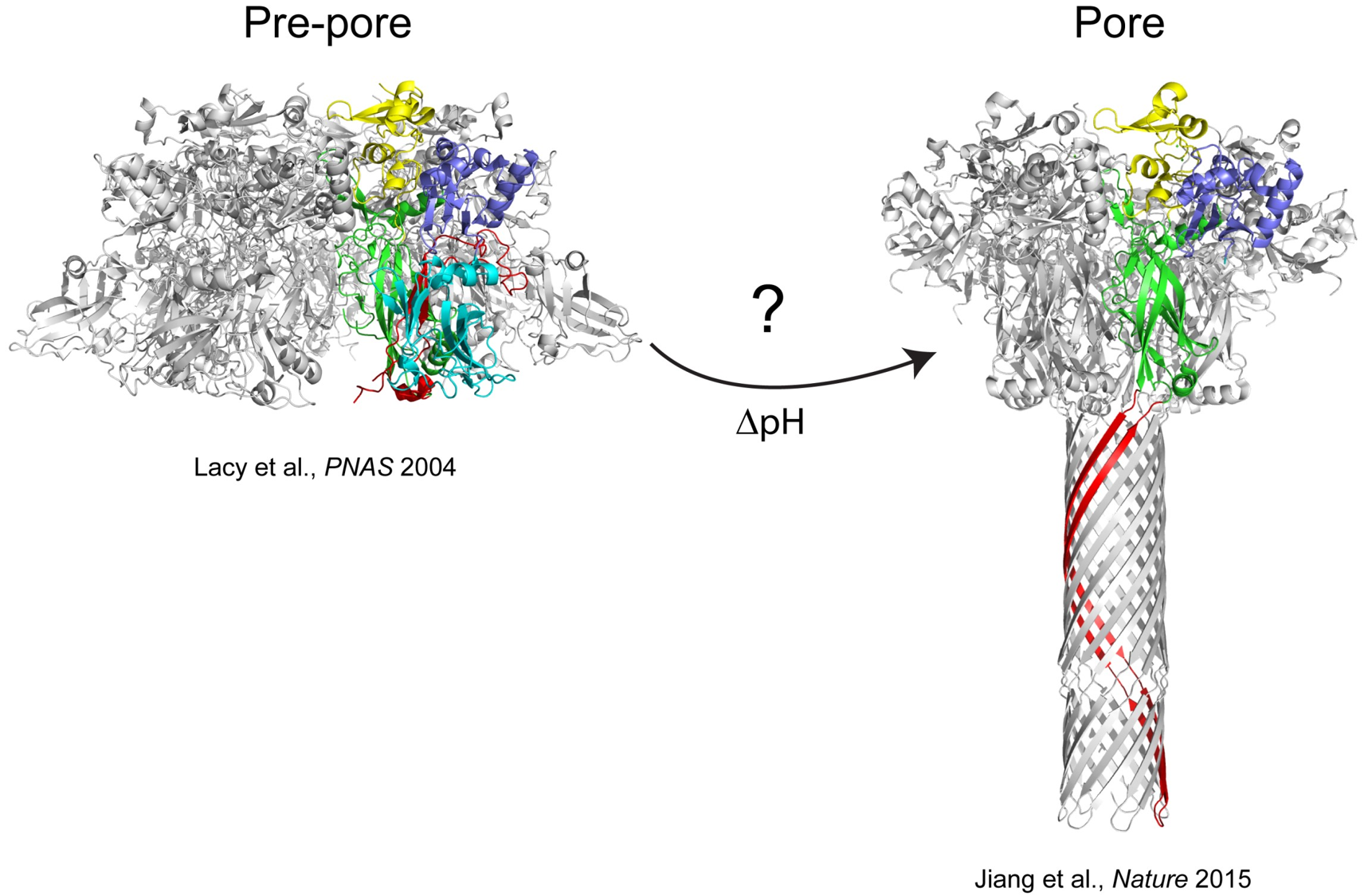
Although the exact mechanism of pore formation is not understood, seminal work from the Collier laboratory at Harvard has shown that pore formation can be prevented by mutating certain amino acid residues within PA (dominant-negative mutants, Sellman et al., JBC 2001). We are currently investigating how these and other mutations prevent pore formation, which may provide clues as to the pathway(s) needed for this large structural change.
The methods we use include circular dichroism spectroscopy, fluorescence (equilibrium and stopped-flow) and fluorine NMR. We have also made use of mutagenesis as well as incorporation of unnatural amino acids to investigate the mechanism of pore formation. In the latter case for instance, we biosynthetically incorporated into PA 2-fluorohistidine, which has a significantly lower pKa than histidine (pKa of ~1). Biosynthetic incorporation does not prevent pore formation of PA unless the soluble form of the host cellular receptor capillary morphogenesis protein 2 (sCMG2) is bound(1). This suggested that histidine protonation could be a trigger for pore formation in the context of the receptor bound state, and lead us to investigate further differences in structure between the receptor bound and unbound states(2).
In collaboration with Masaru Miyagi (Case Western Reserve), we have shown that the thermodynamic stability of PA is greatly increased if sCMG2 is bound (2, 3), and occurs throughout the protein structure. Give the increased stability of PA, and the fact that it is a well-studied immunogen, we are also pursuing studies directed at understanding the role of protein stability in immunogenicity.
Our laboratory has developed methods to biosynthetically incorporate fluorinated amino acids into proteins, and a major focus of our laboratory is to expand the repertoire of fluorinated amino acids that have unique functionalities, and to incorporate these amino acids into proteins for structural studies. Fluorine (19F), is only slightly larger than hydrogen, is a spin ½ nucleus, is 100% naturally abundant and is almost as sensitive as 1H. The NMR spectrum of a fluorinated protein typically exhibits well resolved resonances which are sensitive to environment, allowing the researcher to follow structurally changes in select regions of the protein at a residue-specific level. However, the origin of the fluorine chemical shift in the context of proteins is not well understood. In collaboration with the Mitchell-Koch laboratory, we are investigating the origin of fluorine chemical shifts beginning with simple amino acids and progressing from there to peptides and proteins.
- CHEM 212: General Chemistry II
- CHEM 661: Principles of Biochemistry
- CHEM 662: Biochemistry I
- CHEM 663: Biochemistry II
- CHEM 664: Biochemistry Laboratory
- Halimeh W, Ogunnaike K, Tonelli M, Bann JG, Fast folding of anthrax lethal factor N-terminal domain independent of proline isomerization, Biophysical Journal (2026), doi: https://doi.org/10.1016/j.bpj.2026.04.013.
- Bann, J.G. 2012. Anthrax toxin protective antigen: insights into molecular switching from prepore to pore. Protein Science, 21, 1-12.
- Andra, K.K., Bullinger, J.C., Bann, J.G., and Eichhorn, D.M. 2010. 2-Fluoro-L-histidine. Acta Crystallogr Sect E Struct Rep Online. 66, 2713.
- Wimalasena, D.S., Janowiak, B.E., Lovell, S., Miyagi, M., Sun, J., Zhou, H., Hajduch, J., Pooput, C., Kirk, K.L., Battaile, K.P., and Bann, J.G. 2010. Evidence that histidine protonation of receptor-bound anthrax protective antigen is a trigger for pore formation. Biochemistry 49, 6973-6983.
- Williams, A..S., Lovell, S.R., Anbanandam, A., El-Chami, R., Bann, J.G. (2009). Domain 4 of the anthrax protective antigen maintains structure and binding to the host receptor CMG2 at low pH. Protein Sci. 18, 2277-2286.
- Thomas, C.A., Talaty, E.R., and Bann, J.G. (2009). 3S-fluoroproline as a probe to monitor proline isomerization during protein folding by 19F-NMR. Chem. Comm. 23, 3366-3368.
- Rajapaksha, M., Eichler, J.F., Hajduch, J., Anderson, D.E., Kirk, K.L., and Bann, J.G. (2009). Monitoring anthrax toxin receptor dissociation from the protective antigen by NMR. Protein Sci. 18, 17-23.

